I rarely every come across publicly available datasets that I find interesting from a socio-economic welfare perspective, so when I spotted the Costa Rican Household Poverty (CRHP) dataset on Kaggle, I jumped at the chance to dig in and explore it a bit. Unfortunately, the full training and test sets have still to be released on the site, so in this post, I re-purpose the available data, clean and pre-process it. In Part II of this post, I use this data to evaluate a handful of widely-known classification algorithms using caret.
About the data
This dataset was released on Kaggle as part of a competition sponsored by the Inter-american Development Bank (IDB), with the hopes that the Kaggle community would be able to help them discover new and improved methods for identifying the segments of the population in most need of social assistance.
Just glancing at the data on the Kaggle the page, we can see that there’s a fair amount of information available, with the bulk of the dataset already split into a train set (\(9,557\) x \(143\)) and a test set (\(23,856\) x \(142\)). Oddly, the test set contains 70% of the data, and has one less column, which in this case, happens to be the target column. For the remaining columns, there are two identifiers, plus 140 different columns describing individual and household attributes. I’m not going to go through all of these feature columns here, but the organizers have made a codebook available, and there is a nice summary of all the feature variables on the website.
The core data fields
The main ‘target’ variable in this dataset is a poverty-level classification with four values: “extreme poverty”, “moderate poverty”, “vulnerable”, and “non-vulnerable”.
There are two variables that serve as identifiers–‘Id’ and ‘idhogar’, a unique individual-level identifier and an identifier for which individuals belong to the same household, respectively. An additional variable, ‘parentesco1’ is a dummy that indicates whether a given individual is the head of a household.
Re-splitting the data
The main problem with this dataset is that the available test set does not include the target variable. This was done so that, separately, competing teams could build their models and make their predictions, and organizers could see which teams had the most accurate predictions and select the competition winners. The competition concluded in September of 2018, and the organizers have, at present, still not released the full test set including the target variable, and thus, this data is unusable to us for supervised learning approaches.
The good news is that the training set that is available still contains over 9,500 observations and the target variable, which will allow us to train some basic machine learning algorithms after we re-split it into new training and test sets.
The prime consideration when re-splitting this data is how to manage the two identifiers–‘Id’ and ‘idhogar’. As the goal of the original Kaggle competition was to predict household poverty levels, the ‘idhogar’ grouping is quite important, and indeed the original Kaggle dataset does not divide any households between training and test sets, but rather, keeps the households together on their respective partitions, which makes sense.
While it is quite simple to partition the original training data into new training and test sets using the built-in functions in caret or caTools, neither of these packages currently includes an easy way to partition based on a target variable, while preserving entire household groupings in a balanced way between training and test sets.
In the end, in order to do a proper blocking of the ‘idhogar’ variable and ensure that households were not randomly divided over both the new training and test sets, I performed the data partitioning manually, using dplyr to do repeated random splits of the data in such a way as to preserve the household groupings, and then identifying the seed used where the random split resulted in the most balanced Target distribution in both the new training and test set.
First, we can have a look a the distribution of the ‘Target’ variable in the original CRHP training set:
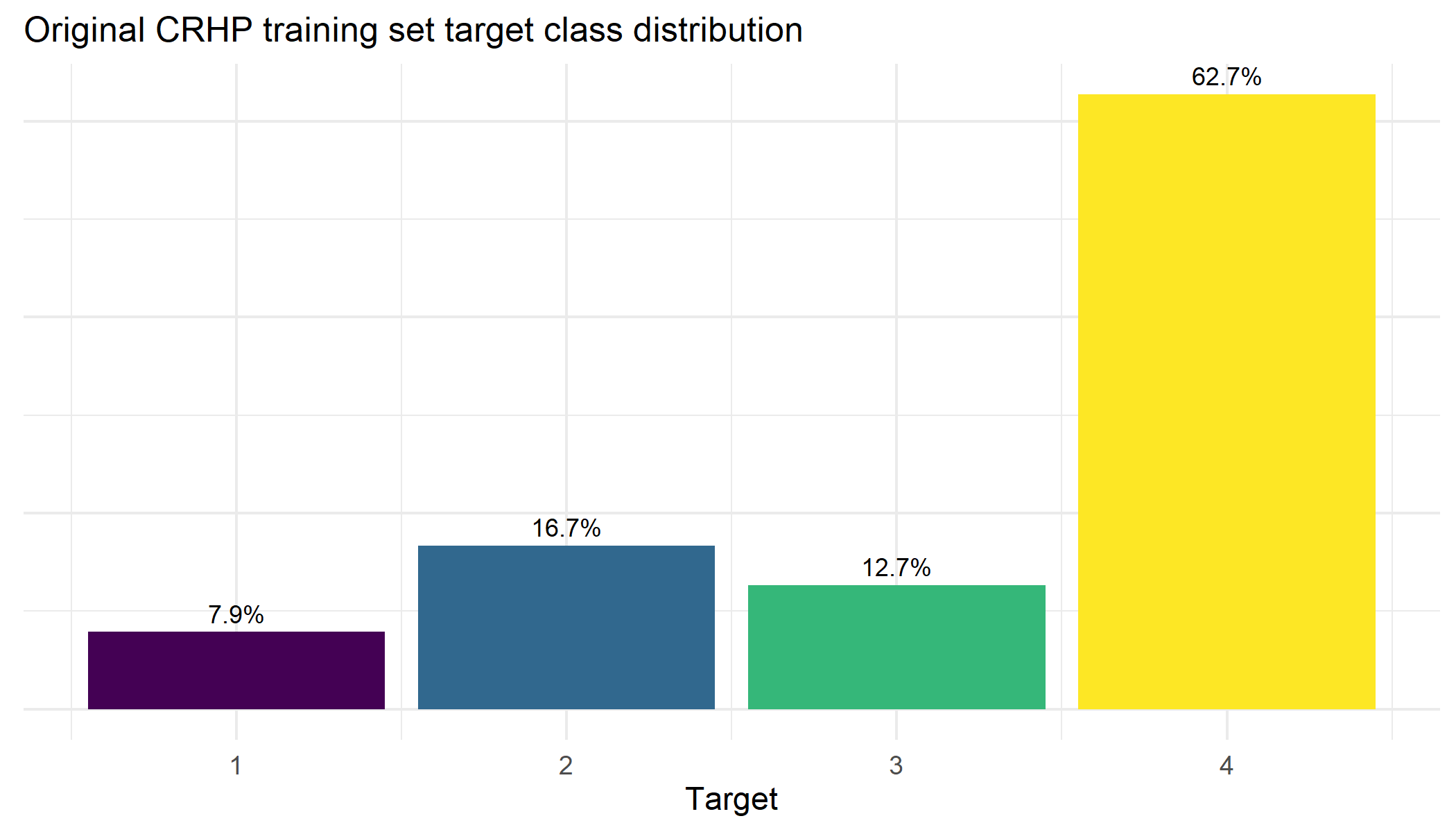
Next, I did 1000 random splits, and then looked for the seed where the frequencies of the different target variable classes between the two sets were the most similar. We can see in the plot below that the balance of the ‘target’ variable classes in our final re-split data is both very well balanced and also matches quite nicely with the original CHRP training set:
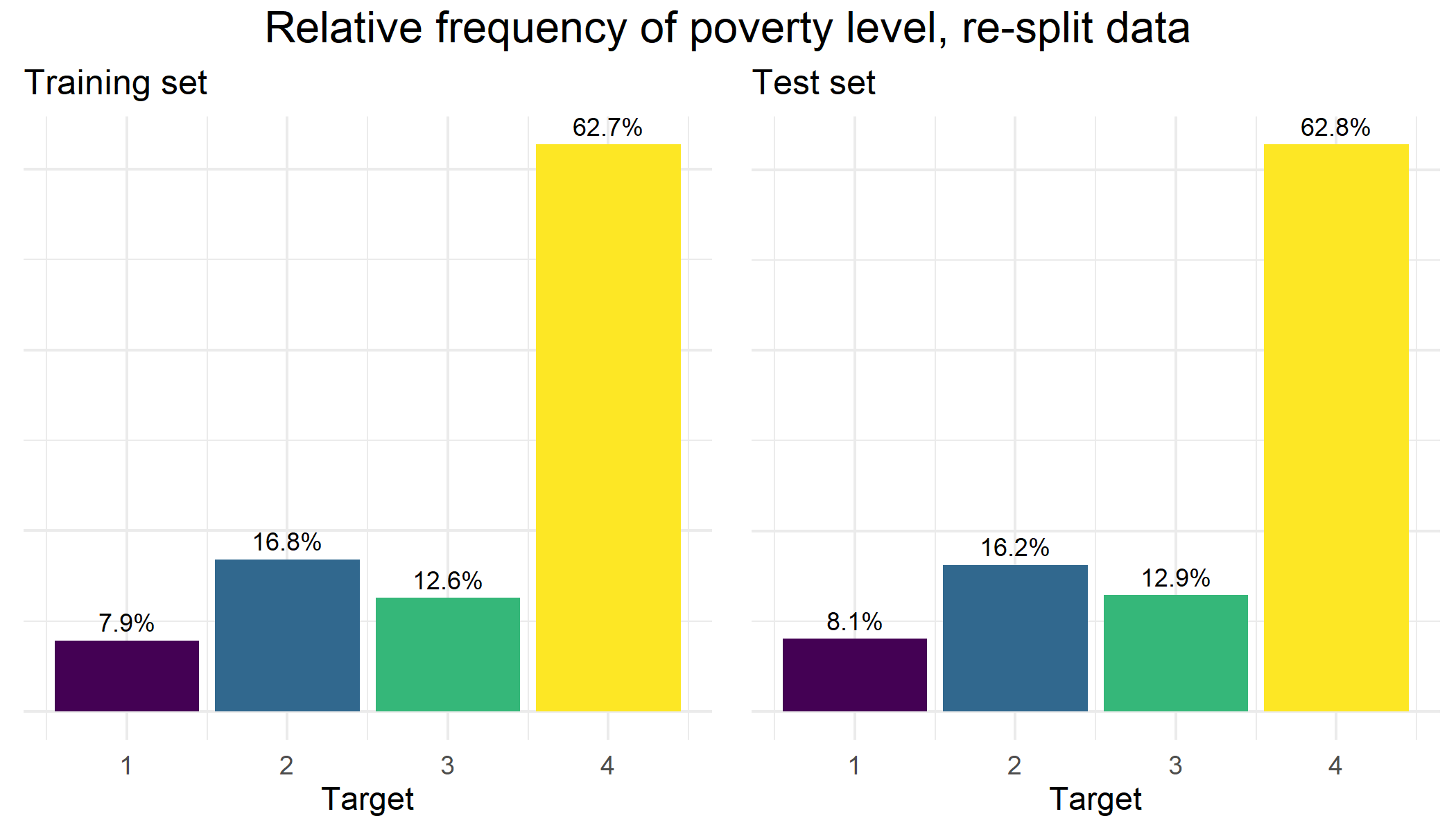
While I do not exploit the household grouping information for this particular analysis, our newly split data captures all of this information and can be used for future projects. Visit my github site to download the CRHP re-split dataset.
Similarly, I also have an Rmarkdown document that contains further analysis on the re-split CRHP data, where I show that the proportion of unique households and the size of households between both sets is also well-balanced.
Cleaning and pre-processing the training data
Now that we have our working dataset, we can quickly go through some basic cleaning and pre-processing. As my goal for this particular analysis is to evaluate the performance of several common classification algorithms, I skip feature engineering completely. As we will see later, our predictive accuracy across all models is pretty low, so this would be a good starting place to re-visit in the future should we wish to find easy way to boost performance.
When we read in our data, we can see that our new train set has 1,908 observations and 143 variables. Furthermore, it looks like the overwhelming majority of this data is already one-hot encoded, and that there are only a handful of other columns that take integer values greater than 1, or that contain character strings:
## Id v2a1 hacdor rooms hacapo v14a refrig v18q v18q1 r4h1
## 1 ID_279628684 190000 0 3 0 1 1 0 NA 0
## 2 ID_f29eb3ddd 135000 0 4 0 1 1 1 1 0
## 3 ID_68de51c94 NA 0 8 0 1 1 0 NA 0
## 4 ID_d671db89c 180000 0 5 0 1 1 1 1 0
## 5 ID_d56d6f5f5 180000 0 5 0 1 1 1 1 0
## 6 ID_ec05b1a7b 180000 0 5 0 1 1 1 1 0
## 7 ID_e9e0c1100 180000 0 5 0 1 1 1 1 0
## 8 ID_3e04e571e 130000 1 2 0 1 1 0 NA 0
## 9 ID_1284f8aad 130000 1 2 0 1 1 0 NA 0
## 10 ID_51f52fdd2 130000 1 2 0 1 1 0 NA 0Missing values
We can check to see what percentage of the dataset is made up of missing values:
# calculate percentage of data that is NA
round(sum(is.na(train))/sum(!is.na(train)) * 100, 2)## [1] 1.64While there are some missing values in our data, the good news is that their percentage is fairly low at 1.66%.
Next we can calculate the percentage of missing values by column to see if there are some variables that are more problematic than others:
na_vars <- colMeans(is.na(train))
na_vars <- na_vars[na_vars > 0]
na_vars <- sort(na_vars, decreasing=TRUE)
na_vars## rez_esc v18q1 v2a1 meaneduc SQBmeaned
## 0.8276898941 0.7665054255 0.7169564649 0.0006536802 0.0006536802Luckily, we can see that there are only five columns in the entire dataset that have missing values, and only three of these have NAs of any meaningful percentage. These variables are:
rez_esc, Years behind in schoolv18q1, number of tablets household ownsv2a1, Monthly rent paymentmeaneduc, average years of education for adults (18+)SQBmeaned, square of the mean years of education of adults (>=18) in the household
Ordinarily, it would be good to dig deeper into these variables to understand why they have so many NAs and see if we can fill them in, but again, as our main interest is fitting multiple models to see how they compare, we will just drop these variables from our analysis entirely as they are reasonably approximated by other variables in the dataset. Similarly, as we continue pre-processing our data, we will just drop other potentially problematic variables straight away rather than spending extra time during this stage to re-work them.
And finally, because we will eventually have to drop these same variables later in our test est, we will create a vector to store the names of all variables we eliminate as we go along:
library(dplyr)
# store names of NA variables to drop
drop_cols <- train %>% select(rez_esc, v2a1, v18q1, meaneduc, SQBmeaned) %>% names() %>% as_tibble()
# drop NA values
train_filtered <- train[ ,!names(train) %in% drop_cols$value]
dim(train_filtered)## [1] 7649 138Zero-variance predictors
Next, we can continue to pare down our feature set by looking for zero-variance or near-zero-variance predictors. As described in the caretdocumentation:
In some situations, the data generating mechanism can create predictors that only have a single unique value (i.e. a “zero-variance predictor”). For many models (excluding tree-based models), this may cause the model to crash or the fit to be unstable. Similarly, predictors might have only a handful of unique values that occur with very low frequencies…
The concern here that these predictors may become zero-variance predictors when the data are split into cross-validation/bootstrap sub-samples or that a few samples may have an undue influence on the model. These “near-zero-variance” predictors may need to be identified and eliminated prior to modeling.
For this, we can use the nearZeroVar function in caret that makes this task fairly simple:
library(caret)
# identify NZV predictors
nzv <- nearZeroVar(train_filtered)
# show names of NZV predictors
train_filtered[, nzv] %>% names()## [1] "hacdor" "hacapo" "v14a"
## [4] "refrig" "pareddes" "paredzinc"
## [7] "paredfibras" "paredother" "pisoother"
## [10] "pisonatur" "pisonotiene" "techozinc"
## [13] "techoentrepiso" "techocane" "techootro"
## [16] "abastaguadentro" "abastaguafuera" "abastaguano"
## [19] "planpri" "noelec" "sanitario1"
## [22] "sanitario5" "sanitario6" "energcocinar1"
## [25] "energcocinar4" "elimbasu2" "elimbasu4"
## [28] "elimbasu5" "elimbasu6" "estadocivil4"
## [31] "estadocivil6" "parentesco4" "parentesco5"
## [34] "parentesco7" "parentesco8" "parentesco9"
## [37] "parentesco10" "parentesco11" "parentesco12"
## [40] "instlevel6" "instlevel7" "instlevel9"
## [43] "tipovivi4" "mobilephone"Again, ideally, we would dig into these variables a bit more to understand what is going on here, but to keep this simple, and as our main identification and outcome variables are not included in this list, we will just drop all 44 of these nzv predictors outright. The procedure is as before:
# store names of NZV variables to drop
drop_cols <- train_filtered[, nzv] %>% names() %>% as_tibble() %>% rbind(drop_cols, .)
# drop NZV variables
train_filtered <- train_filtered[, -nzv]
dim(train_filtered)## [1] 7649 94Linear Combinations
The caret package includes the findLinearCombos function, a simple solution for identifying and removing linearly dependent variables from our dataset. This will further help us reduce dimensionality and improve the performance of our model.
# re-run previous code to convert latest train_filtered back to matrix form
l <- lapply(train_filtered, function(X) as.numeric(factor(X, levels=unique(X))))
m <- as.matrix(data.frame(l))
# find columns to eliminate linear dependencies
comboInfo <- findLinearCombos(m)
# print column names to remove
colnames(m)[comboInfo$remove]## [1] "etecho3" "eviv3" "lugar6" "SQBhogar_nin"When we look up the descriptions of these four variables, we can see that they all belong to sets of dummy variables that have not had one dropped, so they would have perfect multicolinearity if not removed. We will add these four names to our list, and then drop them from train_filtered:
# Store names of linearly dependent variables
drop_cols <- colnames(m)[comboInfo$remove] %>% as_tibble() %>% rbind(drop_cols, .)
# Drop linearly dependent variables
train_filtered <-train_filtered[ ,!names(train_filtered) %in% drop_cols$value]
dim(train_filtered)## [1] 7649 79String variables
As we noted when we first glimpsed at our training data, our data is currently has two variable class types–string variables and numeric variables. Here’s how it looks at the moment:
table(sapply(train_filtered, class))##
## character numeric
## 5 74We have five variables that contain character strings. As string variables needed to be treated differently than numeric ones, it is worth looking into these in more detail before we proceed any further. Here they are:
train_filtered[sapply(train_filtered, is.character)] %>% glimpse()## Observations: 7,649
## Variables: 5
## $ Id <chr> "ID_279628684", "ID_f29eb3ddd", "ID_68de51c94", "ID...
## $ idhogar <chr> "21eb7fcc1", "0e5d7a658", "2c7317ea8", "2b58d945f",...
## $ dependency <chr> "no", "8", "8", "yes", "yes", "yes", "yes", "yes", ...
## $ edjefe <chr> "10", "12", "no", "11", "11", "11", "11", "9", "9",...
## $ edjefa <chr> "no", "no", "11", "no", "no", "no", "no", "no", "no...The first two variables, Id and idhogar, are unique identifiers for individuals and households respectively.
# reorder `idhogar` to second column after `Id`
train_filtered <- train_filtered %>%
select(Id, idhogar, everything())For the other three, which contain a mix of strings and numbers, we can pull up their descriptions from the codebook:
dependency, Dependency rate, calculated = (number of members of the household younger than 19 or older than 64)/(number of member of household between 19 and 64)edjefe, years of education of male head of household, based on the interaction of escolari (years of education), head of household and gender, yes=1 and no=0edjefa, years of education of female head of household, based on the interaction of escolari (years of education), head of household and gender, yes=1 and no=0
We can see all three are derived variables, and the description specifies how they were calculated. As it is not completely clear to me from these descriptions why these variables take an integer, a “yes” or a “no” value, we can select the relevant columns used for the calculations of these variables and see if we can figure out what is going on after a visual inspection.
dependency
To inspect the dependency variable, we select the relevant columns from its calculation and have a look with the following code:
dep_check <- train_filtered %>% select(Id, idhogar, tamhog, tamviv, age, dependency)
head(dep_check, 10)I’m only showing the head of this check for illustrative purposes, but going through the entire subset we just took, the calculations still don’t make sense to me with regards to why a value takes a number, a “yes”, or a “no”, so I will drop the column from our training set for the time being.
drop_cols <- train_filtered %>% select(dependency) %>% names() %>% as_tibble() %>% rbind(drop_cols, .)
# drop NA values
train_filtered <- train_filtered[ ,!names(train_filtered) %in% drop_cols$value]
dim(train_filtered)## [1] 7649 78edjefe and edjefa
Similar to what we did above, we can also subset our data to do a visual inspection of these two variables:
edjef_check <- train_filtered %>% select(Id, idhogar, parentesco1, male, escolari , edjefe, edjefa)
head(edjef_check, 10)Here, it seems that each variable just takes on the value of the escolari variable for whoever is identified as the head of household (parentesco1) for any given group with the same idhogar identifier. When escolari is 1 year, it is being converted to a “yes”, and when it is 0 years, it is being converted to a “no”. We can easily fix this by just converting all yes/no values to 1/0 with the following code:
# replace no/yes with 0/1, convert columns to numeric
train_filtered$edjefe <- as.numeric(ifelse(train_filtered$edjefe == "no", 0, ifelse(train_filtered$edjefe == "yes", 1, train_filtered$edjefe)))
train_filtered$edjefa <- as.numeric(ifelse(train_filtered$edjefa == "no", 0, ifelse(train_filtered$edjefa == "yes", 1, train_filtered$edjefa)))Pre-processing the test set
dim(train_filtered)## [1] 7649 78To recap: at the end our initial pre-processing work, we’ve filtered our original training data down from 143 variables to 78. We’ve eliminated all of the missing values, near-zero-variance predictors, highly redundant variables and linearly dependent variables. We’ve fixed or removed all string variables so that the only two that remain are our individual and household identifiers.
Now all that is left to do for this stage of our project is to apply all of these same changes to our test set to ensure the two will match up:
# drop columns from test set
test_filtered <- test[ ,!names(test) %in% drop_cols$value]
# reorder `idhogar` to second column in test set
test_filtered <- test_filtered %>%
select(Id, idhogar, everything())
# replace no/yes with 0/1, convert columns to numeric in test set
test_filtered$edjefe <- as.numeric(ifelse(test_filtered$edjefe == "no", 0, ifelse(test_filtered$edjefe == "yes", 1, test_filtered$edjefe)))
test_filtered$edjefa <- as.numeric(ifelse(test_filtered$edjefa == "no", 0, ifelse(test_filtered$edjefa == "yes", 1, test_filtered$edjefa)))With that, we’ve taken the original train set from Kaggle, re-split it into new training and test sets, and then cleaned and pre-processed both sets so that they are ready for model fitting and analysis. In the second part of this post, we will use this data to see how some common classification algorithms compare with one another using caret.
